Categories Products for Mass Spectrometry In-vivo Mouse SILAC Proteomics
In-vivo Mouse SILAC Proteomics
In vitro SILAC is a well-established approach for quantification of proteins in cell culture.[1] The same concept can also be applied to living organisms (in-vivo SILAC). The latter approach allows determination of protein patterns of all tissues of an organism with respect to a reference organism. This approach has been successfully applied to the analysis of mouse organs. [2]
Prerequisite for the application of the in-vivo SILAC approach is the availability of an appropriate SILAC diet in which all lysins are replaced by 13C-lysine.
Silantes provides
- SILAC Mouse Diets (diets for other model organisms such as fly and worm are available on request),
- SILAC Mouse Tissues wet and
- SILAC Mouse Tissues lyophilized.
Silantes SILAC Diets for in-vivo labelling of mice[3]
Mice are fed with Silantes SILAC mouse feed according to the scheme in Figure 1 for about 90 days requiring about 800 g Silantes mice feed.
13C-lysine labelled mice feed has been developed in cooperation with the group of Prof. Matthias Mann, Max-Planck-Institute of Biochemistry as a kit containing labelled and unlabelled mouse feed. The kit consists of 13C-lysine-labelled “heavy” diet (B) and unlabelled “light” diet (A). The feed is an artificial, amino acid-based feed using Harlan components.
After the metabolic labelling (feeding) of the mice according to the scheme in Figure 1, the mice are sacrificed. Differences in the protein patterns are determined in analogy to the established SILAC approach in cell culture (see literature for Silantes in vitro SILAC). Instead of 13C-lysine labelled feed, other isotopic formulations are also available on request, e.g. 13C,15N-labelled feed.
A variation of the in-vivo SILAC-approach is the “spike-in” approach.
SILAC spike-in using Silantes 13C6-Lysine-labelled mouse tissue
Figure 1 shows the SILAC spike-in workflow: Differences in protein patterns of unlabelled tissue (A) with respect to unlabelled tissue (B) can be quantified by “spiking-in” a 13C-labelled reference tissue (R). This 13C-lysine reference tissue can be obtained from Silantes directly, as frozen material or lyophilized.
Short outline of the procedure:
The isotopically labelled “heavy” reference tissue (R) is mixed with the unlabelled “light” tissues (A) and (B), respectively. The proteomes of the (A)(R)-mix and (B)(R)-mix are isolated, digested and subjected to LC-MS as shown in the workflow. Therefore, the two peptide amount ratios (A/R) and (B/R) can be determined.
Calculating the ratio (A/R) : (B/R) cancels out the reference amount (R) and yields the ratio of peptides (A/B). The stable isotopically labelled mouse tissue (R) is used as a standard, permitting normalizing (A) with respect to (B), accounting for differences in the isolation procedure in the mixtures (A/R) and (B/R) without affecting the peptide ratios (A/B). (Strain (R) must not necessarily be the same as strain (A) and (B).)
Conclusion: The customer is free in regard to the choise of strain used for the experiment but can use the Silantes 13C-lysine labelled mouse tissue as a reference which is spiked into tissue (A) and (B).
The in-vivo SILAC Lyo-mouse kit
Silantes has recently developed a cost-efficient alternative to the 13C-lysine(6) labelled spike-in wet reference in the form of a 13C-lysine(6) labelled lyophilized reference. This improves the handling of the reference and is more convenient for quantifying protein patterns of single mouse organs.
The lyophilized reference is a ready-to-use sample. It contains 13C-lysine labelled tissue from the respective mouse organ, such as liver, brain, etc. The sample contains about 100 µg of protein which is sufficient for about 3 experiments. The proteins are extracted either with SDS or urea.
- Application note for the in-vivo SILAC Lyo-Mouse buffered in SDS
- Application note for the in-vivo SILAC Lyo-Mouse buffered in SDS (diluted)
- Application note for the in-vivo SILAC Lyo-Mouse buffered in urea
- Application note for the in-vivo SILAC Lyo-Mouse buffered in urea (diluted)
Improved handling with unchanged high quality:
Silantes conducted a comparative study of wet and lyophilized references in cooperation with Prof. Marcus Krüger from Max Planck Institute for Heart and Lung Research.
The results show that the wet sample can be replaced by the lyophilized sample without loss of accuracy.
Figure 2 shows the analysis of the SDS gel patterns of wet liver (STD) and lyophilized liver (Lyo).
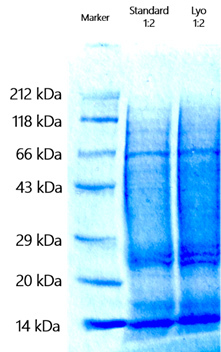
Figure 2: SDS-PAGE of liver tissue samples: Standard (wet liver) and Lyo. (lyophilized liver)
To verify that lyophilization does not change the protein content with respect to the wet sample, both samples were analyzed by mass spectrometry and their peptide patterns were compared.
Please find the results of this analysis below:
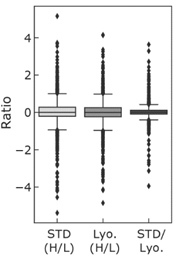
Figure 3: Ratio of labelled (H) to unlabelled (L) peptides of the STD and Lyo.
Figure 3: Row (1) shows the ratio of 13C-labelled (H) to unllabeled (L) peptides of the wet sample (STD); Row (2) shows the same for the lyophilized tissue (Lyo.) and row (3) shows the direct comparison of “STD.” and “Lyo.”.
A comparison of the data in row (1) and row (2) shows that the peptide distributions of “STD” and “Lyo.” have a high degree of similarity. An even greater similarity is observed if wet tissue and lyophilized tissue are directly compared in row (3).
Figure 4 shows a correlation plot using the same data as in Figure 3.
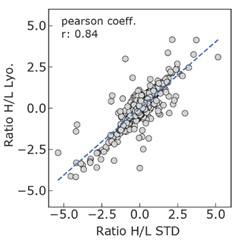
Figure 4: Correlation plot of labelled (H) to unlabelled (L) peptides of the STD and Lyo.
The pearson correlation coefficient of 0.83 indicates a high level of correlation between the peptide data sets from the wet tissue (STD) and the lyophilized tissue (Lyo.), supporting the view that lyophilized tissue is a suitable alternative to the wet tissue.
Figure 5 shows the correlation with the same samples as above, namely the wet liver (STD) and the lyophilized liver (Lyo.), including a new unrelated liver sample (#2) in wet and lyophilized form.
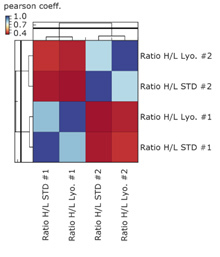
Figure 5: Correlation plot including a new liver sample in wet and lyophilized form
As expected, the data show a lower correlation of #1 with respect to #2 indicating that differences between unrelated liver samples can be detected.
[1] Ong, S.E., Blagoev, B., Kratchmarova, I., Kristensen, D.B., Steen, H., Pandey A., and Mann, M. (2002). Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics, Mol.Cell. Proteomics 1, 376–386.
[2] Ong, S.E., and Mann, M. (2006). A practical recipe for stable isotope labelling by amino acids in cell culture (SILAC). Nat. Protoc. 1, 2650–2660.
[3] Krueger M., Moser M., Ussar S., Thievessen I., Luber Ch.A, Forner F., Schmidt S., Zanivan S., Faessler R. and Mann M.(2008). SILAC Mouse for Quantitative Proteomics Uncovers Kindlin-3 as an Essential Factor for Red Blood Cell Function, Cell 134, 353–364.
SILAC Mouse Diet
|
|
SILAC Mouse Tissue lyophilized
|



